Predicting Cancer Using Support Vector Classifier Model
Machine learning can help identify patterns and relationships in medical data that were previously unknown, allowing for more accurate diagnoses and treatments. This is particularly useful in the early detection of diseases such as cancer, where a robust predictive model can be developed using a support vector classifier to identify the presence of malignant or benign cells.
Early detection is crucial in the medical industry and can potentially save lives. In addition, machine learning can also help in categorizing diseases and identifying the root cause of a disease, making it an important tool in the overall medical diagnosis process.
Key takeaways from this article
We will explore the following questions in detail:
- What methods are used to predict the presence of malignant tumors (cancer cells)?
- What steps are involved in implementing an SVM classifier for predicting breast cancer?
- How can we evaluate our model using the confusion matrix and ROC curve?
- In which medical domains can machine learning be helpful?
- Which major companies are contributing to this area?
- Possible interview questions on this topic.
Let's first define our problem statement in detail and proceed to build a solution.
Problem Understanding
Breast cancer is the most common type of cancer among women and is the second leading cause of cancer deaths in women. It occurs when there is abnormal growth of cells in the breast tissue, also known as a tumor. However, not all tumors are cancerous - they can be benign (not cancerous), pre-malignant (pre-cancerous), or malignant (cancerous). Diagnostic tests such as MRI, mammogram, ultrasound, and biopsy are commonly used to detect breast cancer.
Next, we see the test, which helps us get features about the cell to predict whether it is malignant or benign.
FNA Test
The Fine Needle Aspiration (FNA) test is a quick and simple procedure that involves removing fluid from a part of the body where there is swelling or tenderness. The fluid collected from this test is used to create a labeled dataset that can be used to develop a machine learning model for breast cancer classification. This data includes certain features, along with the results of the FNA test, and is used to determine if a patient has malignant cells that could potentially lead to breast cancer. If a person tests positive, they can take steps to reduce the risk of testing positive in the future.
Machine learning techniques, such as artificial neural networks, gradient boost method, and support vector machines (SVM), can be used in conjunction with clinical data to accurately predict cases. In this article, we will focus on building an SVM model for predicting cancer cells based on specific observations. Let's begin the implementation steps.
Cancer Classification Model Implementation Steps
Step 1: Importing Libraries and Loading the dataset
To begin, we will load several libraries including pyplot for plotting graphs, numpy for handling arrays and interacting with numerical arrays, and pandas for better handling of datasets in the form of data frames. We will use the loadbreastcancer dataset from the sklearn library for this purpose. The breast cancer dataset is a classic and straightforward binary classification dataset that is included with the Scikit-learn library. It can be imported using sklearn.datasets.loadbreastcancer.
from matplotlib import pyplot as plt
import numpy as np
from sklearn.svm import SVC
import pandas as pd
from sklearn.datasets import load_breast_cancer
dat=load_breast_cancer()We have loaded the dataset, but it is essential to understand the content of the data. We see these steps in the next section.
Step 2: Understanding the data
The dataset has a dimension of 569 x 32, with each instance labeled ‘M’ or ‘B,’ where M = malignant and B = benign.
cancer_features=pd.DataFrame(dat.data,columns=dat.feature_names)
'''
Index([‘Unnamed: @", ‘diagnosis’, ‘radius_mean’, ‘texture_mean’,
‘perimeter_mean’, ‘area_mean', ‘smoothness_mean’, “compactness_mean',
‘concavity_mean’, ‘concave points_mean’, ‘symmetry mean’,
“fractal_dimension_mean', ‘radius_se’, ‘texture_se’, ‘perimeter_se’,
‘area_se’, ‘smoothness_se’, ‘compactness_se’, ‘concavity se’,
‘concave points_se’, ‘symmetry se’, ‘fractal dimension_se’,
‘radius_worst", "texture worst’, "perimeter_worst’, "area worst’,
'smoothness worst', ‘compactness worst’, ‘concavity worst’,
concave points worst’, ‘symmetry worst’, ‘fractal_dimension_worst'],
dtype=" object")
'''The above-shown attributes are the features to be used to predict cancer.
Note: The first feature, ‘Unnamed: 0’, is an index and can be excluded from the final features.
We have seen column names, but as the feature vector size is large, it isn’t easy to see the relation between features. So we know the method of Data Visualization in the next section.
Step 3: Data Visualization using RedViz
from pandas.plotting import radviz
radviz(dat.ix[:,1:],"diagnosis",color=['red', 'green'])In this dataset, there is a highly non-linear relation between the features; hence, a robust classifier is needed to make any prediction based on it. We have used RadViz ( a non-linear multi-dimensional visualization library) to visualize the dataset of every feature.
Introduction to RedViz library
RadViz is a technique that allows us to visualize N-dimensional data in a 2D space. In RadViz, each dimension of the dataset is represented as an anchor, evenly spaced around a unit circle. Each line in the data corresponds to a point in the projection, connected to each dimensional anchor by a spring. It's important to note that each dimension is normalized before being plotted on the RadViz plot. The closer the linked columns are to each other on an optimized plot, the more correlated they are. The (x, y) coordinate corresponding to a particular feature vector will change if the order of columns is changed.
In the image, each instance in the dataset is represented as either "red" or "green" according to the labels. The visualization shows a clear correlation between the instances in the dataset, making it necessary for a strong classifier to solve this problem. While we have visualized the data, it cannot be directly input into the model and needs to be pre-processed, which we will cover in the next section.
Step 4: Data Preprocessing for loadbreastcancer dataset
This step involves several activities, such as:
- Assigning numerical values to categorical data (target labels): To define the target values in this task, we can use a label encoder. The label encoder can be imported using the command, from sklearn.preprocessing import LabelEncoder. Once it is imported, we can create an instance of the label encoder and fit it to the target attribute column (diagnosis).
li_classes = [dat.target_names[1], dat.target_names[0]]
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
target_encoded = pd.Series(dat.target)
target = le.fit_transform(target_encoded)-
Standardize every instance of the features: In this step, data standardization can be performed, which will orient the data and a zero mean and unit standard deviation.
X_new = (X - mean(X))/Standard_deviation(X)
cancer_features=cancer_features.drop(['mean perimeter','mean area','mean radius','mean compactness'],axis=1)
STD=StandardScaler()
cancer_features=STD.fit_transform(cancer_features)Now we want to train the model. We see in the next section how to train a model using SVM.
Step 5: Model Formation
Kernelized support vector machines (SVMs) are powerful methods for mapping a nonlinear dataset to a relatively linear space to classify instances of the dataset. Therefore, we will use SVM for this task to achieve better performance.
SVM is a machine learning algorithm that tries to draw a hyperplane to separate nonlinear data features so that we can distinguish between different data labels and perform classification. The creation of the hyperplane involves many steps behind the scenes.
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score- Obtaining the training and testing data sample: We can use the traintestsplit module from sklearn.model_selection library to divide the above dataset into training and testing sets. The split can be done in a ratio of 75:25.
x_train,x_test,y_train,y_test=train_test_split(cancer_features,target,test_size=0.25,random_state=0)- Creating an instance of the model: Import the support vector classifier (SVC) from the SVM module in the sklearn library using from sklearn.svm import SVC and create an instance of the model.
model=SVC(C=1.2,kernel='rbf')
model.fit(x_train,y_train)
y_pred=model.predict(x_test)Step 6: Overall Pipeline creation
A pipeline allows you to group and execute components sequentially. It can be imported from the sklearn library using from sklearn.pipeline import make_pipeline. This pipeline will process the input by sequentially applying the components. The figure below shows that the training dataset is fed into the pipeline. Once the dataset is standardized, it is passed to the support vector classifier (SVC) to solve a classification problem.
We can use the previously prepared training set and use the .fit method to fit the training data on the classifier pipeline.
The above figure shows the model pipeline to demonstrate the flow. The model used is SVC, which has a lot of tunable parameters, like:
- Regularizer, C: This parameter has a default value of 1. It is a positive float value that inversely relates the strength of the regularization to the value.
- Kernel: The kernel transforms the data into a different form to make it easier for the classifier to classify. The most commonly used kernel is the radial basis function (rbf) kernel because it can account for nonlinear relationships in the data.
The parameters discussed above are important to consider when building the model. Other parameters, such as the gamma value, can be set to auto to allow the model to adjust them internally.
We have created a model, but we need to evaluate it to determine whether it is a good fit for the data. We will examine this in the next section.
Step 7: Performance Evaluation of the SVM model
We have solved a classification problem so that the model can be evaluated on several classification evaluation metrics.
print("accuracy: ", accuracy_score(y_test, y_pred))
print("precision: ", precision_score(y_test, y_pred))
print("recall: ", recall_score(y_test, y_pred))
print("f1: ", f1_score(y_test, y_pred))
print("area under curve (auc): ", roc_auc_score(y_test, y_pred))Accuracy Score
This score is simply the percentage of correct prediction in the test set. For the above-given configuration, the accuracy is close to 95.8%.
Confusion Matrix
The confusion matrix can be imported from the metrics module of the sklearn library. It is used to compare the predicted output of the test set with the ground truth. You may be wondering why we need to use a confusion matrix instead of just using accuracy directly. The confusion matrix allows us to predict precision, accuracy, and f1 score, which can provide a more detailed evaluation of the data. You can read it in detail in this blog.
ROC
ROC is the plot between False Positive and True Positive in the plot. It can be imported using. from sklearn.metrics import roc_curve.
We have successfully built our model and it is performing well. Now let's examine some other medical fields where machine learning could be beneficial.
Different domains in medical science where machine learning can help
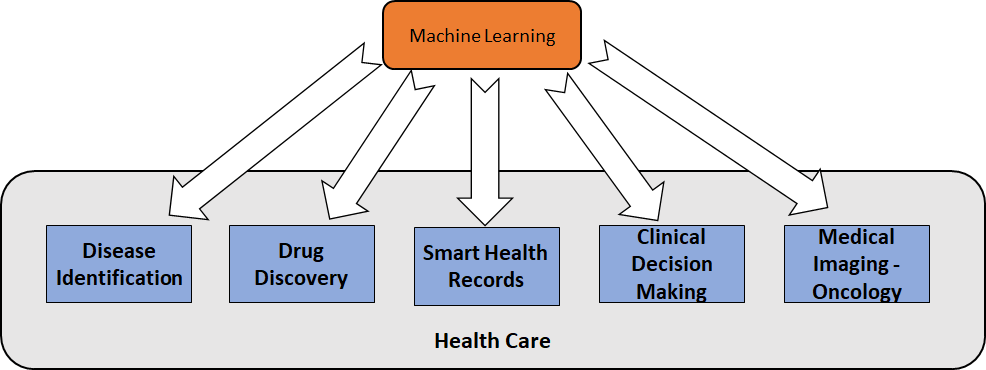
There are many applications of machine learning in the medical field, ranging from disease diagnosis to advanced image processing techniques that aid in tasks previously only performed by pathologists and microbiologists. Here are five significant areas in the medical domain where machine learning is making a significant contribution:
- Disease identification
- Drug discovery
- Smart health records
- Clinical decision making
- Medical imaging in oncology
Case studies of Companies Use-case
Early diagnosis of cancer is a crucial step in saving lives. With this in mind, many multinational companies (MNCs) have invested significant time and resources into this area. Let's examine some of the work being done by these MNCs.
DeepMind by Google
Google has used machine learning approaches to predict different forms of cancer, with a particular focus on lung cancer, which is the leading cause of death, even more than breast cancer. Their algorithm has outperformed radiologists in identifying cancer cases from CT scan diagnostic images.
IBM Watson & Mayo Clinic
IBM and Mayo Clinic have teamed up to use IBM's cognitive computing research capabilities, including artificial intelligence, computer vision, and natural language processing, to enhance the diagnosis of cancerous tissue and complement human expertise in the clinical field.
Mayo Clinic provides clinical trial data, while IBM's technology helps extract information more quickly and accurately than a doctor could, identifying patients who are best suited for Mayo Clinic's clinical trial criteria. This includes genomic analysis, matching patients to appropriate clinical trials, and generating evidence to support formal treatment recommendations as the standard of care. Watson Health, offered by IBM, is an example of these cognitive services.
Possible Interview Questions
If you are planning to include this project in your resume, you may be asked the following questions in a machine learning interview:
- What are classification problems?
- Why did you choose SVM? What other algorithms could you have used instead of SVM?
- What features were included in the final feature set?
- Is it possible to convert this to a multi-class classification problem?
- What additional pre-processing could be done on the final data?
- What are type-1 and type-2 errors?
Conclusion
Machine learning is playing a crucial role in saving lives by helping doctors identify diseases quickly and providing possible treatments. Breast cancer is a common disease that affects women and causes more than 4 million deaths annually, accounting for 14% of all cancer deaths. Algorithms like SVM and artificial neural networks (ANNs) are capable of detecting the possibility of breast cancer in patients, which could take doctors a long time to identify.
Early detection can greatly reduce the death rate from breast cancer. Many companies, including Google and IBM, have invested in cancer identification. Additionally, there are also many health apps, platforms, and smart watches that track people's health in order to promote a healthier future.
Next blog: Introduction to KNN Algorithm in Machine Learning
Enjoy Learning, Enjoy Thinking!
Share Your Insights
More from EnjoyAlgorithms
Self-paced Courses and Blogs
Coding Interview
OOP Concepts
Our Newsletter
Subscribe to get well designed content on data structure and algorithms, machine learning, system design, object orientd programming and math.
©2023 Code Algorithms Pvt. Ltd.
All rights reserved.
